Four experts with over five years of MR image-quality assessment expertise.
Blinded Comparison:
Raters performed side-by-side comparisons of images with and without PMC.
Image presentations were randomised to avoid bias.
Scoring System:
Image quality rated from 1 (worst) to 10 (best) based on motion artefacts.
Objective Image Quality Assessment
Evaluation Metrics
Average Edge Strength (AES): Measures the average strength of edges in an image, indicating the clarity and sharpness of structural details. (higher AES -> higher image quality)
Gradient Entropy (GE): Evaluates the randomness and complexity of the gradient distribution, reflecting the overall texture and contrast of the image.
(lower GE -> higher image quality)
High-Resolution MRI:
PMC significantly improves image quality in high-resolution scans.
Only three scenarios showed statistically significant improvement, but overall subjective and objective measures indicate better quality with PMC.
Applicability of PMC:
PMC is beneficial for high-resolution scans even in the absence of deliberate motion.
Effective for healthy, compliant participants.
Image Quality Assessment (IQA)
Limits for Subjective IQA: Variability Among Observers, Bias, Limited Reproducibility, Time-Consuming, Lack of Objectivity, ...
Current no-reference IQA metrics are not always robust and reliable for MR images
Study: SSIM Prediction for Detection and Quantification of Motion Artefacts in Brain MR Images
[Structural Similarity Index Measures: SSIM]
Collection of motion artefact-free MR scans (from 1.5 to 7.0T)
With/Without Data Augmentation (brightness/contrast adjustment)
Artificial corruption of MR images
SSIM calculation between corrupted and motion-free images
Training ResNet models (ResNet-18 and ResNet-101) to predict SSIM using only corrupted images as input and the calculated SSIM value
Testing models on both artificially corrupted and clinical data
SSIM Predictions and Model Evaluation
Results:
Best model: ResNet-18 trained with data augmentation
Worst model: ResNet-101 trained without data augmentation
The analysis of clinical data samples revealed that the highest agreement rate between subjective scores and SSIM predictions was 77.7% for ResNet-101 trained with data augmentation.
This work is under review:
IEEE Access, "Automated SSIM Regression for Detection and Quantification of Motion Artefacts in Brain MR Images"
Preprint: arXiv:2206.06725
What does it happen when the extra hardware required by PMC is not available?
(Like for example in a clinical setting.)
Solution: We can use Deep Learning based Retrospective Motion Correction!
Study: Retrospective Motion Correction of MR Images using Prior-Assisted Deep Learning
Collection of 100 motion artefact-free MR scans from IXI public dataset
Artificial corruption of MR images
Image Priors (Similar Slices):
10 similar slices (same position, same contrast) from different subjects.
using T1 and PD-weighted images from the same subject.
motion correction only for T2-weighted images.
Network Architectures
Baselines: ReconResNet* and U-Net**.
Prior Supply Techniques:
Multi-Channel Network: Concatenated motion-corrupted image with priors.
Dual-Branch Network: Main branch for corrupted image, auxiliary branch for priors.
*Chatterjee, Soumick, et al. "Reconresnet: Regularised residual learning for mr image reconstruction of undersampled cartesian and radial data." Computers in biology and medicine 143 (2022): 105321.
**Ronneberger, Olaf, et al. "U-net: Convolutional networks for biomedical image segmentation." MICCAI 2015
Some result
Conclusion
Key Findings:
Additional contrast images from the same subject are more beneficial than similar slices from different subjects.
Multi-channel and dual-branch approaches showed improvements, especially for ReconResNet.
Medical Imaging Workshop of 34th Conference on Neural Information Processing Systems (NeurIPS 2020), Vancouver, Canada
Study: Generalised Retrospective Motion Correction (RMC) using Deep Learning and Contrast Augmentation
Collection of 600 motion artefact-free MR scans from IXI public dataset, in-house scans @3.0 and 7.0T
Data Augmentation (brightness/contrast adjustment)
Artificial corruption of MR images
No image priors required
Training of ReconResNet model*
*Chatterjee, Soumick, et al. "Reconresnet: Regularised residual learning for mr image reconstruction of undersampled cartesian and radial data." Computers in biology and medicine 143 (2022): 105321.
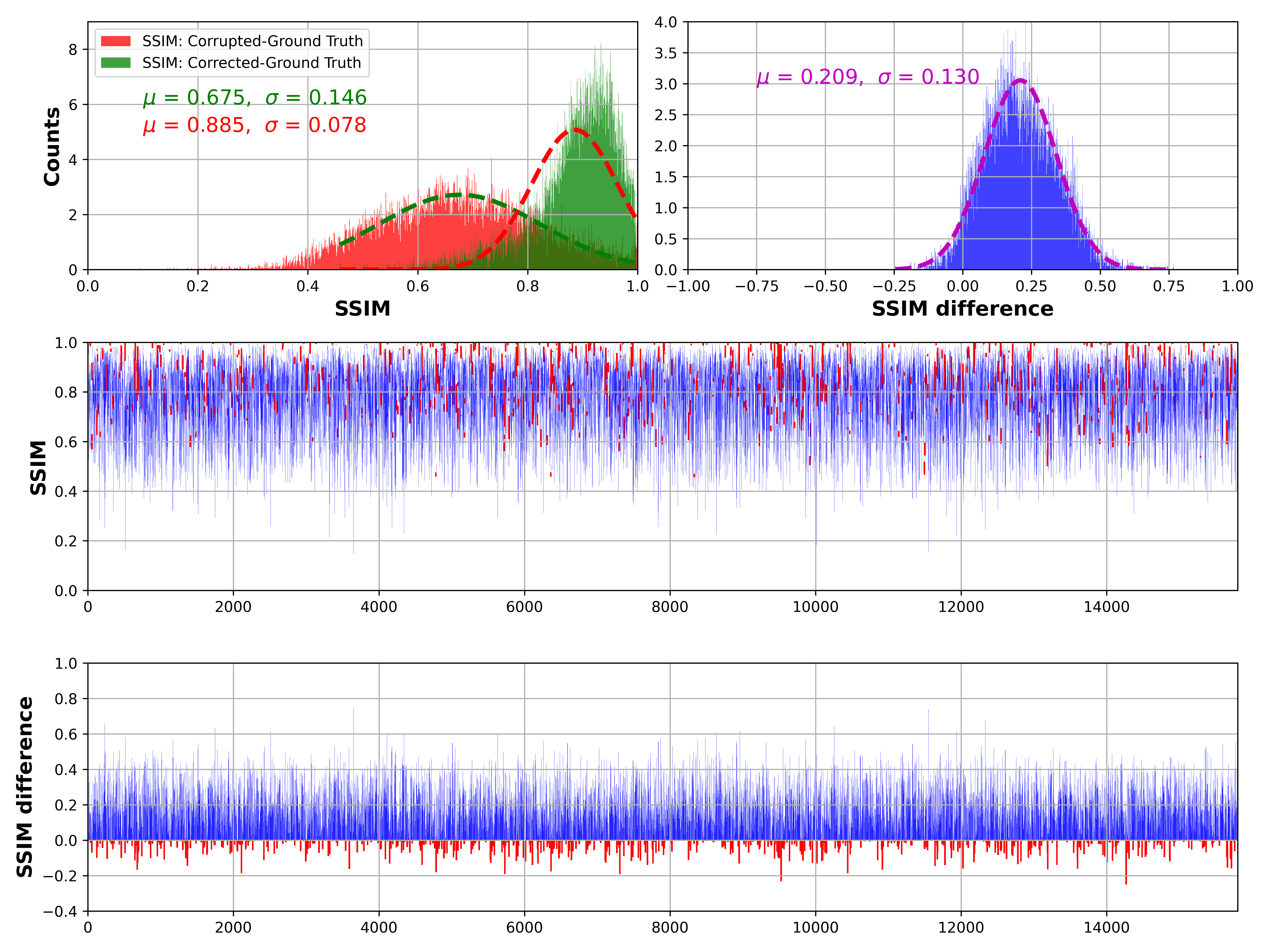
Results
Average SSIM improvement from 0.7 to 0.9.
Consistent performance across different experimental runs.
Qualitative results show significant enhancement in image quality.
Conclusion
Proposed method demonstrates broad generalisation across various MRI contrasts and artefacts.
Notable improvements in image quality and robustness.
Manuscript under preparation
Some test on clinical data!
Conclusions
Demonstrated the effectiveness of PMC
Presented a novel method for motion artefacts evaluation
Novel Deep Learning methods for RMC
Using image priors
Using generalised correction models
Future work
Amalgamation of PMC and RMC with DL
Motion model approach
PMC, then DL-based RMC
A big thank you to:
Prof. Oliver Speck,
Kathrin Schulze, Frank Godenschweger,
Daniel Stucht, Hendrik Mattern, Astrid Wollrab,
Yi-Hang Tung, Mahsa Fatahi, Peter Schulze,
Sebastian Hupfeld, Renat Yakupov, Oleg Posnansky,
Urte Kägebein, Yan Arnold, Nicolas Huch, Michel Pohl,
Uten Yarach, Weiqiang Dou, Shan Yang, Myung-Ho In,
Dominik Kolmann, Tobias Leutritz, Denis Kokorin,
Tino-Johannes Lüttge, Falk Lüsebrink-Rindsland
Prof. Steffen Oeltze-Jafra, Yannic Waerzeggers,
Max Dünnwald, Juliane Müller
Special thanks to Soumick Chatterjee and Max Dünnwald
Funding information
Initial Training Network, funded by the
FP7 Marie Curie Actions of the European
Commission (FP7- PEOPLE- 2012-
ITN- 316716) and the National Institutes
of Health (1R01-DA021146), the DFG
(DFG- MA 9235/1-1), and the federal state
of Saxony-Anhalt (“I 88”)
- **Affine Transformations**: Aligns images with real-time adjustments for consistency.
effectiveness varies by sequence and motion type.
However, these studies focused on a single contrast
and included only small cohorts (1-11 subjects). Thus, a
comprehensive assessment of the performance of PMC for
high- resolution MRI in a larger cohort is missing. To that
end, we acquired data of 21 healthy subjects for four different
sequences at 7 T. To mimic a routine research setting, sub-
jects were asked not to move (unlike most motion- correction
studies that correct for intentional motion). The images were
assessed subjectively through expert ratings and objectively
with image metrics. The aim of this work was to verify and
quantify whether PMC can significantly improve image qual-
ity at 7 T, for healthy compliant subjects and in the regime of
“quasi-no- motion” as introduced by Stucht et al.1 This study
is not meant to assess the performance of PMC for a broader
population of subjects inexperienced with MRI
- 21 healthy volunteers were scanned over the course of two independent 75-minute long sessions
(14 males, 31.5±6.1 years old, and 7 females, 27.3±3.4 years old).
- $T_1$, $T_2$, $PD$ and $T_2^*$-weighted images for each subject were acquired
+ Scores reflect the degree of corruption due to motion.
### Statistical Analysis
- **Intraclass Correlation Coefficient (ICC)**:
+ Calculated using Pingouin to assess agreement among raters.
+ Majority of images with PMC ON showed high or very high quality.
- **Evaluation Metrics**:
+ Five out of six groups showed higher image quality with PMC.
+ One group showed inconsistent results between subjective and objective metrics.
Influence of Context, Inability to Capture All Aspects, Training and Expertise Required.
do not consistently apply to all MR acquisition types
, indicating a need for more robust evaluation methods.
**To address this gap, a novel approach has been developed that leverages deep learning for SSIM prediction, providing a more reliable and accurate assessment of image quality.**
The extra-hardware required by PMC is not available in a clinical setup
## BUT WE CAN USE DEEP LEARNING TO TRY TO SOLVE THIS PROBLEM
- The RMC (Retrospective Motion Correction) approach utilizes deep learning to address the challenges of PMC.
- Trained and Tested deep learning models that can correct motion artefacts in MRI images retrospectively:
+ **Retrospective Motion Correction of MR Images using Prior-Assisted Deep Learning**
+ **Generalised Retrospective Motion Correction (RMC) using Deep Learning and Contrast Augmentation**
## Data Preparation
- **Dataset**: 100 participants’ T1, T2, and PD images from IXI Dataset.
- **Artificial Motion Corruption** as done before
- Modified TorchIO’s RandomMotion transformation.
- Simulated movements: rotation from -1.75 to +1.75 degrees (no translation).
## Image Priors
- **Similar Slices**:
- 10 similar slices (same position, same contrast) from different subjects.
- Used for motion correction of T2-weighted images.
<!-- - **Different Contrasts**:
- Utilized T1 and PD images from the same subject as priors for motion-corrupted T2 images.
*S. Chatterjee et al. A deep learning approach for reconstruction of undersampled cartesian and radial data. In ESMRMB 2019, Oct. 2019.
- **Dataset**: 100 participants’ T1, T2, and PD images from IXI Dataset.
- **Artificial Motion Corruption** as done before
<!-- - Modified TorchIO’s RandomMotion transformation.
- Simulated movements: rotation from -1.75 to +1.75 degrees (no translation).
- Image Priors (**Similar Slices**):
- 10 similar slices (same position, same contrast) from different subjects.
- Used for motion correction of T2-weighted images.
- **Different Contrasts**:
- Utilized T1 and PD images from the same subject as priors for motion-corrupted T2 images.
## Network Architectures
- **Baselines**: Modified ReconResNet and U-Net.
- **Prior Supply Techniques**:
- **Multi-Channel Network**: Concatenated motion-corrupted image with priors.
- **Dual-Branch Network**: Main branch for corrupted image, auxiliary branch for priors.
## Results
- Similar slices did not improve motion correction.
- Different contrasts significantly enhanced motion correction.
- Multi-channel and dual-branch approaches outperformed ReconResNet.
- Only multi-channel strategy significantly improved U-Net.
- **Future Work**:
- Explore dual-branch approach and impact of skip connections.
- Expand dataset and introduce different motion corruption types for robustness.
Chatterjee, Soumick, et al. "Reconresnet: Regularised residual learning for mr image reconstruction of undersampled cartesian and radial data." Computers in biology and medicine 143 (2022): 105321.
+ 64 feature maps, 56 residual blocks, and PReLU activation.
+ Perceptual loss function using a pretrained ResNeXt 101 model.
- This extension introduces a deep learning method for RMC in MRI using:
- ReconResNet model.
- Novel contrast augmentation and artificial motion corruption techniques.
-
---
## Methods
### Data
- Collected from 3T and 7T MRI Siemens scanners.
- **Dataset**: 600 training, 160 validation, and 158 testing image volumes.
- Only slices containing brain tissues were considered.
### Data Processing
- Random slice selection from 3D volumes.
- Noise removal and Min-Max normalization.
- Padding and resizing to 256x256.
- Contrast augmentation (as for the SSIM prediction).
### Motion Corruption (as for the SSIM prediction)
- Two artificial motion corruption techniques:
1. **TorchIO Functions**: Random ghosting and motion.
2. **In-House Method**: Simulates real-world motion corruption using random parameters.
### Model and Training
- Deeper version of ReconResNet with:
- 64 feature maps, 56 residual blocks, and PReLU activation.
- Perceptual loss function using a pretrained ResNeXt 101 model.
- Optimized using Adam with a learning rate of 3x10⁻⁴ for 2000 epochs.
---
+ Motion model approach
+ PACE with SVR
+ PMC with IRS
+ PMC, then DL-based RMC
---
# An extraordinary thank you to:
<div class="two-columns">
<div class="column">
<strong>Irene Bartolini (my wife)</strong>,
<br>
<strong>Agnese & Benedetta (my twin daughters)</strong>,
<br>
<strong>and all my family and friends for their invaluable help and support</strong>
</div>
<div class="column">
</div>
</div>